Do you know which type of animal cell is the most variable in terms of its shape and size?
It's the sperm cell, or spermatozoa, which undergoes rapid changes and evolution. A biology and genomics project at the Natural History Museum (UiO) seeks to unravel why and how sperm cells have diversified within the diverse group of passerines, or songbirds.
To do this, they are awarded 150,000 CPU hours on Saga, the preferred supercomputer among many biology users. The Natural History Museum has collected extensive data on sperm length in various bird species over the past 15 years.
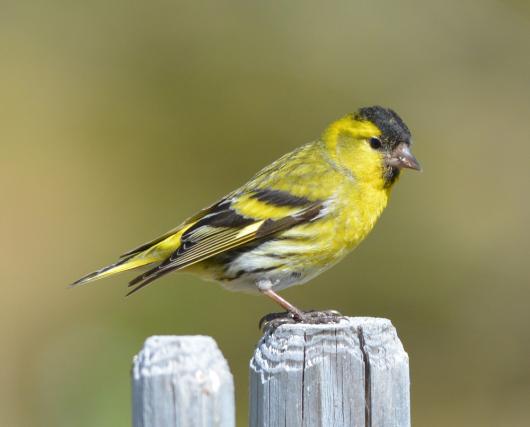

Genetic and evolutionary dynamics of passerine sperm variation
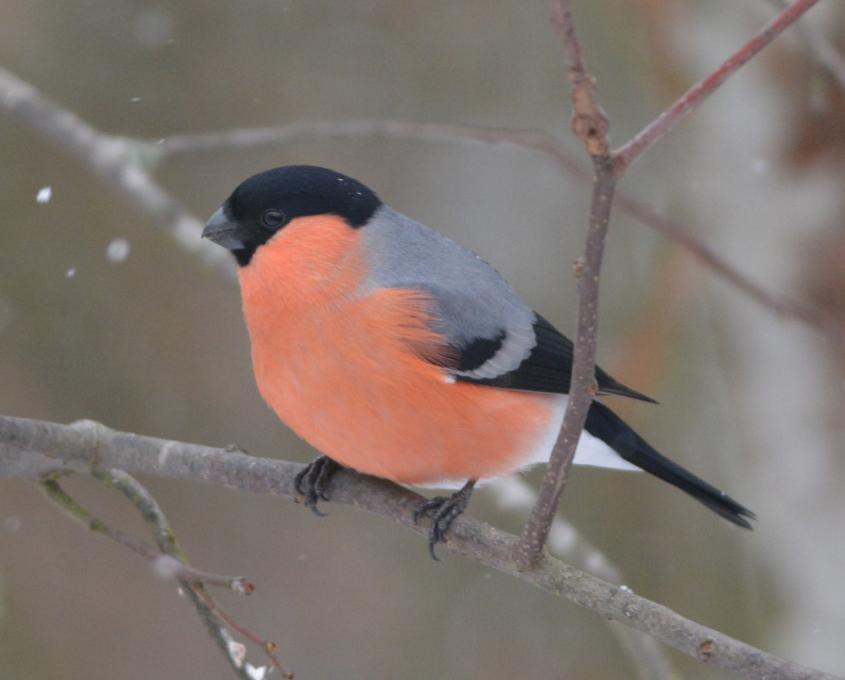

This data is invaluable for studying how bird sperm has evolved in terms of its structure and function over evolutionary history. To better understand the genetics behind sperm variation, the museum collected blood and tissue samples along with sperm samples, enabling researchers to study the genes responsible for these differences more effectively. The project will employ various scientific methods to identify which genes contribute to the variation in sperm length among passerine birds. Additionally, the project has expanded to explore the proteins in sperm, seminal fluid, and egg yolk membranes.
Sperm cells are evolving, even though their main role in fertilisation remains crucial. These cells are influenced by two opposing forces: one that ensures they perform their essential job in fertilisation and another that drives changes in their traits to improve their chances of fertilising eggs. These variations in sperm traits can affect their success in fertilising eggs, how females choose sperm, and even maintain the separation of different species by hindering interbreeding.
Supercomputing power needed
The researchers need supercomputer resources for routinely dealing with large amounts of genomic or transcriptomic (the study of cellular RNA transcripts) data.
—We are reconstructing phylogenetic relationships of bird species and estimating divergence times which require millions of iterations and often take days to complete. We use these divergence time estimates of species and populations to better understand the speed of sperm morphological evolution. Additionally, we use transcriptomic data from testes (bulk RNAseq and single-cell sequencing) to compare how the genes responsible for sperm morphology have changed across species, says Erica H. Leder, Associate Professor II at the Natural History Museum.
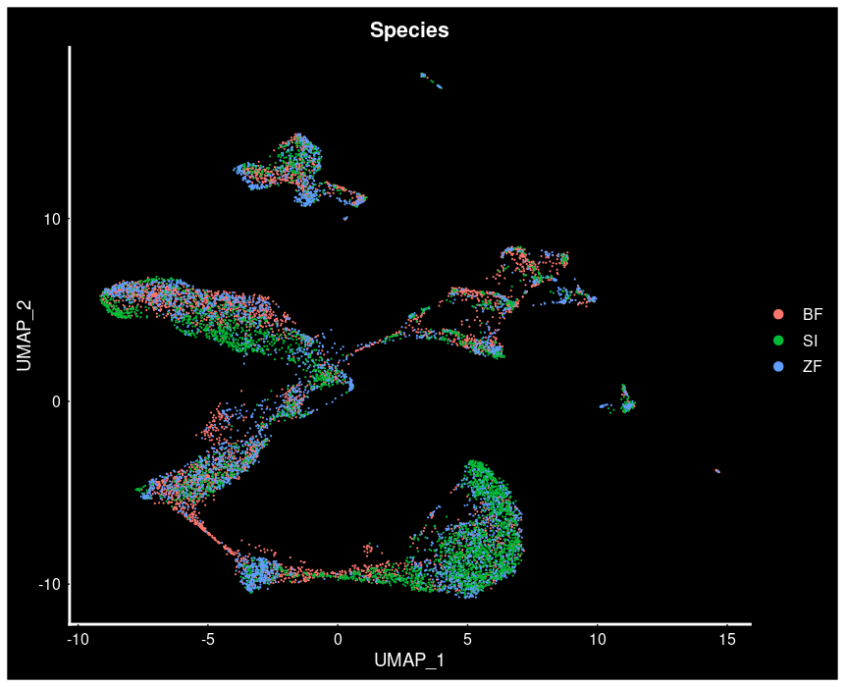
These analyses take advantage of the large capacity for parallelisation and RAM that are available using a supercomputer, in this case, Saga. The main outcome of this project will be a comprehensive list of genes linked to and underlying the differences in sperm structure among passerine birds, and a greater understanding of how sperm morphology contributes to speciation.
The project leader is Postdoctoral Fellow Emma Whittington.